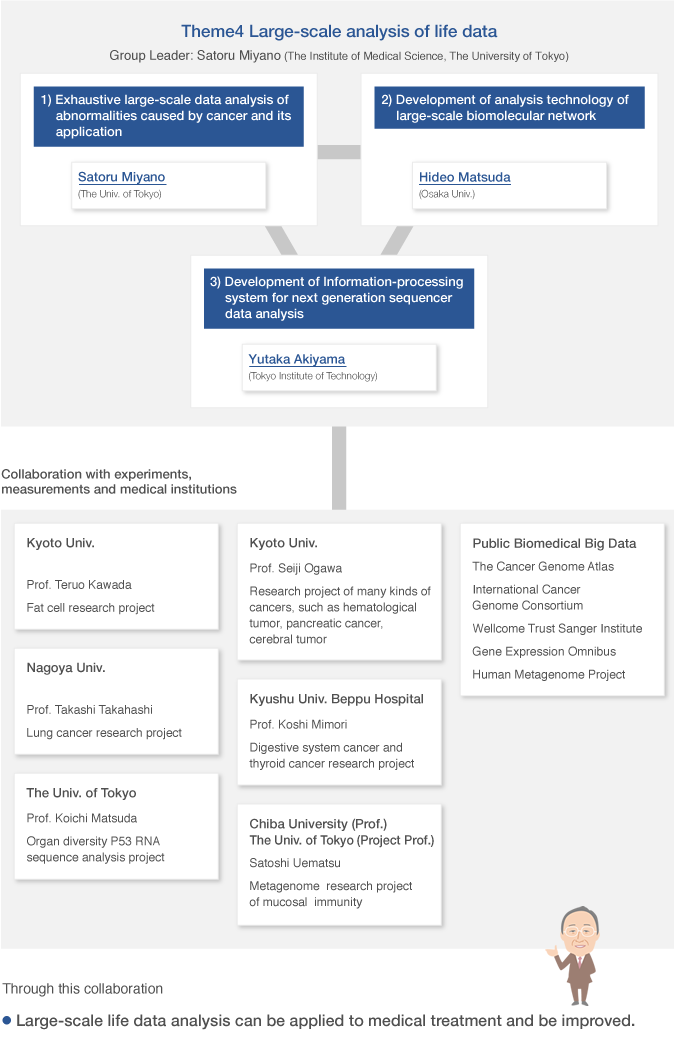
Theme4 Large-scale analysis of life data
Large-scale analysis of life data

Group Leader:
Satoru Miyano
(The Institute of Medical Science, The University of Tokyo)
Summary
Cancer and metabolic syndrome are of the biggest interest in health for the people.
Our high-speed and high-performance analysis of large-scale life data, such as genomic information, contributes to the progress of personalized medicine to tackle cancer and other diseases.
Personalized medicine is a concept that involved providing the most appropriate treatment and health management to respective individuals, based on the utilization of their genomic data and lifestyle-related data. It is expected that patients can get access to safe, reliable and efficient medical care.
In particular, cancer is a systemic disease, where abnormalities in the genome build up infinitely to proliferate, metastasize, and be resistant to anticancer drugs in a very complex manner. In Japan, half of the people are facing cancer somewhere in their lives. At present, researchers around the world are cooperating to analyze abnormalities of genome and gene expression of cancer tissues on a large scale, but the huge amount of data becomes a bottleneck. K computer is used for dramatically accelerating optimal prevention and diagnosis of and therapeutic development for cancer by maximizing the data analysis and exhaustively revealing abnormalities caused by cancer.
Obesity needs medical intervention. It is proved that fat cells of mice in a room at 4 degrees Celsius change to beige anti-metabolic cells to generate heat 100 times as much as that generated in skeletal muscles. Clarification of the mechanism possibly leads to strategies to reduce obesity from a new point of view and decreases the prevalence of obesity and metabolic syndrome dramatically. K computer is very effective to shed light on the mechanism to burn the fat.
Approach during the Past Five Years and its Achievements
Publications
1. R. Araki, S. Seno, Y. Takenaka, H. Matsuda:
“An estimation method for a cellular-state-specific gene regulatory network along tree-structured gene expression profiles“
Gene, Vol.518, No.1, pp.17-25 (2012)
2. Naoki Matsushita, Shigeto Seno, Yoichi Takenaka, Hideo Matsuda:
“Metagenome fragment classification based on multiple motif-occurrence profiles“
PeerJ, Vol.2, e559 (2014)
3. Keisuke Kataoka, Yasunobu Nagata, Akira Kitanaka, Yuichi Shiraishi, et al.:
“Integrated molecular analysis of adult T cell leukemia/lymphoma“
Nature Genetics (2015)
4. Ryutaro Uchi , Yusuke Takahashi, Atsushi Niida, Teppei Shimamura, Hidenari Hirai, Keishi Sugimachi, Genta Sawada, Takeshi Iwaya, Junji Kurashige, Yoshiaki Shinden, Tomohiro Iguchi, Hidetoshi Eguchi, Kenichi Chiba, et al.:
“Integrated Multiregional Analysis Proposing a New Model of Colorectal Cancer Evolution“
PLOS Genetics (2015)
Group organization